URMC / Labs / Ermolenko Lab / Projects / Folding and Dynamics of mRNA
Folding and Dynamics of mRNA
mRNA has an intrinsic propensity to form extensive secondary structure. Structured regions of mRNA are known to regulate mRNA splicing, translation and degradation.
In collaboration with the Mathews lab (University of Rochester), we recently showed that all mRNAs and lncRNAs intrinsically form secondary structures with short end-to-end distances (https://www.ncbi.nlm.nih.gov/pubmed/30337527 ). We now explore biological implications of this structural feature of RNA molecules.
 In collaboration with the Wedekind lab (University of Rochester), we study structural dynamics of riboswitches, structured RNA elements located within the 5’ untranslated regions of bacterial mRNAs that regulate transcription elongation and translation initiation by sensing the presence of various metabolites (https://www.ncbi.nlm.nih.gov/pubmed/30328747 ).
In collaboration with the Wedekind lab (University of Rochester), we study structural dynamics of riboswitches, structured RNA elements located within the 5’ untranslated regions of bacterial mRNAs that regulate transcription elongation and translation initiation by sensing the presence of various metabolites (https://www.ncbi.nlm.nih.gov/pubmed/30328747 ).
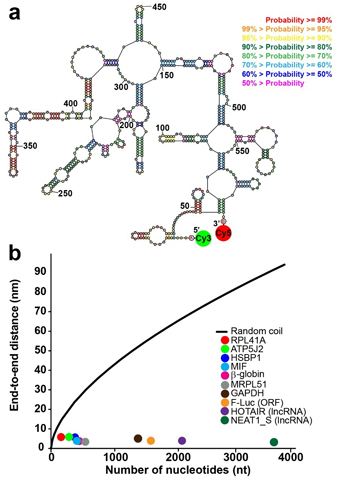
Figure legend: RNA ends are brought within FRET distance via the formation of intramolecular basepairing interactions. (a) An exemplary secondary structure from the ensemble of structures of rabbit β-globin mRNA lacking poly(A) tail predicted by free energy minimization. Base pair probabilities, predicted with a partition function, are color-coded. In order to measure the end-to-end distance by FRET, the 5’ and 3’ ends of mRNA were conjugated with donor (Cy3) and acceptor (Cy5) fluorophores, respectively, as indicated. (b) Average end-to-end distances of mRNAs and lncRNAs, which were folded in the presence of 100 mM KCl and 1 mM MgCl2, were determined by ensemble FRET measurements and plotted as a function of RNA length: yeast RPL41A mRNA (red), firefly luciferase mRNA (orange), rabbit β-globin mRNA (magenta), human ATP5J2 mRNA (green), HSBP1 mRNA (indigo), MIF mRNA (blue), MRPL51 mRNA (grey), GAPDH mRNA (brown), HOTAIR lncRNA (purple), and NEAT1_S lncRNA (dark green). The black line shows theoretically predicted end-to-end distance of unstructured RNA, assuming a freely-jointed chain model (From https://www.ncbi.nlm.nih.gov/pubmed/30337527 ).
« back to all projects