Research Projects
Mechanisms of Nonsense-Mediated mRNA Decay (NMD)
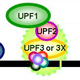 NMD is a splicing- and translation-dependent pathway that targets not only disease-associated transcripts but also naturally occurring transcripts as a way for cells to maintain homeostasis in a changing environmental millieu (reviewed in Maquat et al., 2010, Cell 142:368-74; Hwang and Maquat, 2011, Curr Opin Genet Dev 21:422-30; Popp and Maquat, 2013, Annu Rev Genet 47:139-65; Popp and Maquat, 2014, Mol Cells 37:1-8; Kurosaki and Maquat, 2016, J Cell Sci 129:461-7; Popp and Maquat, 2016, Cell 165:1319-22).
NMD is a splicing- and translation-dependent pathway that targets not only disease-associated transcripts but also naturally occurring transcripts as a way for cells to maintain homeostasis in a changing environmental millieu (reviewed in Maquat et al., 2010, Cell 142:368-74; Hwang and Maquat, 2011, Curr Opin Genet Dev 21:422-30; Popp and Maquat, 2013, Annu Rev Genet 47:139-65; Popp and Maquat, 2014, Mol Cells 37:1-8; Kurosaki and Maquat, 2016, J Cell Sci 129:461-7; Popp and Maquat, 2016, Cell 165:1319-22).
Learn more about Mechanisms of Nonsense-Mediated mRNA Decay (NMD)
Post-transcriptional Gene Regulation by Double-stranded RNA-binding Proteins
 We have found that STAU1, which is a double-stranded RNA binding protein, recruits the NMD factor UPF1 to certain mRNA 3'-untranslated regions (3'UTRs) so as to elicit SMD in a translation-dependent mechanism (reviewed in Park and Maquat, 2013, Wiley Interdiscip Rev RNA 4:423-435). Using microarray analyses, we have identified a number of mRNAs that are naturally down-regulated by SMD.
We have found that STAU1, which is a double-stranded RNA binding protein, recruits the NMD factor UPF1 to certain mRNA 3'-untranslated regions (3'UTRs) so as to elicit SMD in a translation-dependent mechanism (reviewed in Park and Maquat, 2013, Wiley Interdiscip Rev RNA 4:423-435). Using microarray analyses, we have identified a number of mRNAs that are naturally down-regulated by SMD.
Learn more about Post-transcriptional Gene Regulation by Double-stranded RNA-binding Proteins
