URMC / Labs / Rustchenko-Bulgac Lab / Projects / Sorbose-utilizing Mutants Monosomic for Ch5
Sorbose-utilizing Mutants Monosomic for Ch5
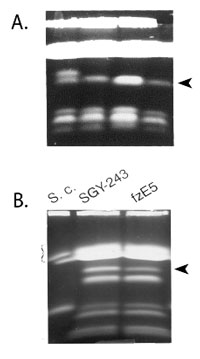
The loss and gain of one homolog
of a chromosome represents a novel
form of gene regulation.
Detailed studies of sorbose-utilizing mutants monosomic for Ch5 provided an understanding of some aspects of the mechanism of regulation as based on diminution of chromosome copy number. Ch5 monosomy and re-duplication (Figure A) occur at high frequencies with a high degree of fidelity, which fulfills a requirement for regulatory systems. Ch5 monosomy induces expression of the metabolic gene for sorbose catabolism, SOU1, which is carried on Ch4. The loss of an entire Ch5 is required because Ch5 carries multiple functionally redundant genes encoding the negative regulators of sorbose utilization that are scattered along the chromosome. We have suggested that the mode of gene regulation involving rearrangements of large specific parts of genome implies that regulatory and metabolic genes in C. albicans are distributed non-randomly over the chromosomes. An entire chromosome thus, can act as a single regulatory unit, a feature not previously considered. Studies of the model system of resistance to sorbose unraveled the unanticipated complexity of regulation by chromosome copy number. Ch5 carries multiple unique Control of Sorbose Utilization (CSU) genes encoding negative regulators of growth on sorbose, however the total number of CSUs remains elusive. Expression of at least two of CSUs is modulated by embedded antisense transcripts. While expression of some genes on monosomic Ch5 is decreased conform diminished gene copy number, expression of some other genes is dosage compensated to the disomic level. We also established a precedent of epigenetic regulation by showing differential increase of H4 acetylation on the monosomic Ch5.
« back to all projects