Computer Simulation Captures Immune Response to Flu
Researchers have successfully tested for the first time a computer simulation of major portions of the body’s immune reaction to influenza type A, with implications for treatment design and preparation ahead of future pandemics, according to work accepted for publication, and posted online, by the Journal of Virology. The new “global” flu model is built out of preexisting, smaller-scale models that capture in mathematical equations millions of simulated interactions between virtual immune cells and viruses.
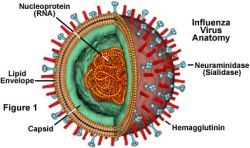
Mathematical models and computer simulations have been used to understand viral infections and immune response to those infections, including influenza type A viruses, which can cause severe human disease. Type A flu viruses are classified by differing versions of key proteins, hemagglutinin (H) and neuraminidase (N), on their outer surfaces that attract attention by the human immune system, but that are always changing. Thus, some forms of both seasonal flu and swine flu are designated H1N1 because of their related, but differing surface proteins. The “bird flu” virus that emerged in 2004 is designated as H5N1. The newest, much-publicized strain of H1N1 swine flu is believed to have caused deaths and hospitalizations because victims’ immune systems did not recognize the latest variations in these surface proteins. Each year, seasonal flu causes approximately 36,000 deaths in this country for the same reason.
A team of immunologists, mathematical modelers, statisticians and software developers created the new model over three years within the Center for Biodefense Immune Modeling at the University of Rochester Medical Center. The project was led by Hulin Wu, Ph.D., principal investigator of the project, director of the Center for Biodefense Immune Modeling (CBIM) and division chief of the Department of Biostatistics and Computational Biology, and by Martin S. Zand, M.D., Ph.D., co-director of the CBIM. The work was funded by the National Institute of Allergy and Infectious Diseases (NIAID), part of the National Institutes of Health, and the U.S. Department of Energy.
“High-speed, accurate computer simulation tools are urgently needed to dissect the relative importance of each attribute of viral strains in their ability to cause disease, and the contribution of each part of the immune system in a successful counterattack,” said Zand. “Real world experiments simply cannot be executed fast enough to investigate so many complex surprises, and we must keep pace with viral evolution to reduce loss of life.”
Small Window
The human immune system is composed of two major branches. The innate system provides immediate protection against bacteria, viruses and other microbes. The second aspect is the more time-consuming, precise and thorough adaptive immune system pumps out vast numbers of immune cells on the hope that one will have the right receptor protein to link up with, and become activated by, any invader encountered. When this occurs, the immune cell expands into an army of clones specifically selected to attack that organism. Workhorses of the adaptive system include “killer” T cells that attack cells that have been infected by viruses before the virus can turn them into virus factories, and helper T cells that orchestrate other parts of the immune response. Both types of T cells are spurred into action by antigen-presenting cells, which engulf the invading microbes and “show” them to the T cells so that the T cells can recognize the infection. In lymph nodes, helper T cells, in turn, cause the second major player in the adaptive response, the B cell, to start mass-producing antibodies, proteins that glom onto and eliminate them from the body.
The new model predicts how rapidly the T and B cells respond to influenza type A virus infection. For the purposes of the simulation, the model set aside the role of the innate immune system, but the researchers plan to add that aspect to future simulations. Depending on the pathogen at hand and a given patient’s past exposure, either T cell or B cell responses may play a larger role in clearing the virus. One cell type may lead the immune counterattack in a person with a first-time infection, and another in a patient who has been infected before or vaccinated.
T cells partner with dendritic cells to make careful decisions about which key pieces of viruses will trigger a full-scale immune counterattack. Dendritic cells roam the body, checking each particle they come across for a self or invader “label.” Upon encountering an invader, a dendritic cell will “swallow it,” cut it up, and carry the pieces to the nearest lymph node. Once there, proteins inside the dendritic cell “present” protein fragments of the virus on the cell’s surface for consideration by T cells gathered there. Once activated by high enough levels of target protein fragment, T cells become capable of destroying the virus.
Using a statistical method called sensitivity analysis, Wu, Zand and colleagues used the new model to compare the individual contribution of various immune cells to clear influenza from the body, including the presentation of fragments of pathogenic proteins on the surface of dendritic cells versus the action of helper and cell-killing T cells versus the direct glomming onto disease-related particles by antibodies. The model predicted that drugs and vaccines that change the presentation by dendritic cells of disease-related protein fragments (antigens) to T cells more strongly impact whether or not a given patient beats the flu than other immune factors in patients infected for the first time. Simulations also suggest that an increase in antigen presentation by dendritic cells can compensate for the increased infection rate of highly virulent influenza strains. Existing adjuvants – drugs that strengthen vaccines but have no effect alone – are available that enhance the ability of dendritic cells to present viral antigens to T cells.
Experiments on the effects of anti-viral drugs on a “flu-infected” virtual immune system also accurately predicted that lowering the viral load or spread within two days of symptoms enables rapid control of the infection. These results match known limitations of neuraminidase inhibitors like the widely stockpiled Tamiflu and Relenza, which inhibit viral reproduction in infected cells, but are only effective if given soon after infection. The model also predicted that antiviral therapies may be more effective if taken in combination, but only if administered within two days of infection. Unlike chronic HIV infection, the acute nature of influenza means there is a narrow time window during early infection when interfering with viral replication can reduce viral load.
“Heterosubtypic immunity” refers to the ability of human immune defenses primed to fight one strain of flu to provide broad protection should other strains of flu be encountered later. Whether or not a single vaccine can protect against any of the many H1N1 Influenza viruses would depend on this property. The team’s model confirmed that heterosubtypic immunity to influenza depends on a specific quality, the number of killer T cells present in the lungs of a given individual at the start of infection. Thus, near-future vaccines against swine flu for instance may work better if designed to evoke local immune memory cells in the lungs, rather than administered through the bloodstream.
In addition, the model argues that a strong antibody response, along with local T cell expansion, will be important to ensure protection against future pandemics. The findings suggest that some unconventional therapies that rapidly boost immune responses might be effective in a worst case scenario of a pandemic influenza virus. One possibility is the treatment of those front line of the epidemic (e.g. healthcare workers) with an injection of the liquid portion of blood (serum) taken from patients who have previously been infected and successfully fought off an influenza virus. This serum would contain antibodies against the virus at hand. Current antiviral treatments only work if given with 48 hours of infection, and vaccines take several weeks to start working, leaving a gap that might be filled by immune sera injection.
Because its genetic makeup is new, little is known about the distribution of preexisting immunity to current H1N1 “swine” flu across populations, the immune response to the virus or the efficacy of available vaccines. The team in Rochester hopes to contribute to the building of models that simulate swine flu infection across the entire U.S. population to better predict its course.
Along with Wu and Zand, major project contributors included Ha Youn Lee, Sung Yong Park, Jeanne Holden-Wiltse and Hongyu Miao in the Department of Biostatistics and Computational Biology at the Medical Center; David Topham, Joseph Hollenbaugh, Tim Mosmann and Brian Ward within the David H. Smith Center for Vaccine Biology & Immunology and the Department of Microbiology and Immunology; and John Treanor and Xia Jin in the Division of Infectious Diseases within the Department of Medicine. Alan Perelson led a partnering effort at the Los Alamos National Laboratory in New Mexico.
“The right computer model can provide a precise, hands-on way of measuring just how good our theories are about how the system responds to pandemic virus, and how to strengthen our defenses,” said Wu.

