Molecular Recognition & Biosensing
Research in the Miller group focuses on two fundamental areas: the control of biomolecular interactions through the synthesis of new small-molecule probes, and the observation of biomolecular interactions through the development of novel optical sensing technologies. In the area of control, we are particularly interested in the sequence-selective recognition of RNA. New RNA sequences with important functions in basic biology and human health and disease are being discovered at an ever-increasing rate, and yet our ability to target these sequences specifically is still at a rudimentary stage. To address this gap, we are applying techniques of molecular design and a novel combinatorial method of small-molecule evolution called Dynamic Combinatorial Chemistry, which allows us to rapidly prototype sequence-selective RNA binding molecules. Thus far we have used this methodology to RNA targets important in Myotonic Dystrophy and HIV. Protein-targeted small-molecule discovery projects are also of interest, and current projects include the mechanism of tight junction formation and the transport of beta-amyloid across the blood-brain barrier.
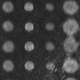
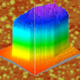
To the end of achieving better methods of observing biomolecular interactions, our group has a longstanding program in the use of the optical properties of nanostructured materials as the basis for new biosensors and diagnostic tools. Two examples of current efforts include Arrayed Imaging Reflectometry (AIR) and sensors based on two-dimensional photonic crystals (2-D PhC). AIR relies on the creation of a near-perfect antireflection coating on the surface of a silicon chip; binding of a biomolecular target destroys this antireflective condition and is visible by a change in reflected light. This allows for highly multiplexed (10's to 1000's of targets) and quantitative detection. Photonic crystal sensors, on the other hand, offer the possibility of ultrasensitive detection: for example, a major long-term goal of our work is the production of sensors that can effectively detect one virus in a blood sample.

Benjamin L. Miller, Ph.D.
Principal Investigator
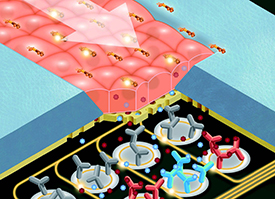 Graduate Student John Cognetti’s Research Featured on Inside-Cover of Lab on a Chip
Graduate Student John Cognetti’s Research Featured on Inside-Cover of Lab on a Chip
A photonic biosensor-integrated tissue chip platform for real-time sensing of lung epithelial inflammatory markers
- Dual-Scale StaphAIR: Predictive Modeling for the Diagnosis of S. aureus Infection via Simultaneous Detection and Quantification of Cytokines and Antibodies.; Analytical chemistry. 2026 May 16.
- A human synovial tendon-on-a-chip models key features of peritendinous adhesions and offers a new approach methodology for testing anti-fibrotic drugs.; bioRxiv : the preprint server for biology. 2026 Apr 07.
- A multi-readout photonic sensor for rapid diagnosis of Von Willebrand disease.; Npj biosensing; Vol 3(1), pp. 7. 2026 Feb 02.
- QuickDraw: detecting HIV in whole blood using an integrated paper-based consumable that enables direct amplification of purified RNA from paper.; Lab on a chip. 2026 Jan 05.
News
Affiliations
- Biochemistry & Biophysics
- Biomedical Engineering
- Dermatology
- Center for AIDS Research
- Center for RNA Biology: From Genome to Therapeutics
- Del Monte Institute for Neuroscience
- NIH T32 Training Grant in Cellular, Biochemical & Molecular Sciences
- NIH T32 Training Grant in Immunology
- Biochemistry & Molecular Biology Ph.D. Program
- Biomedical Engineering Ph.D. Program
- Biophysics, Structural & Computational Biology Ph.D. Program
- Toxicology Ph.D. Program
October 16, 2024
Scientists Developing Microchips with Brain and Lung Tissue to Study Viral Neuroinflammation
January 9, 2024
New NIH-funded Center Could Soon Reduce the Need for Pharmaceutical Trials on Animals
June 15, 2023
Miller Lab Welcomes Undergraduate Sara Boyer
April 12, 2023
Gabrielle Kosoy, Biophysics PhD Candidate in Dr. Ben Miller's Lab, wins 2nd place in UR Three Minute Thesis (3MT) Competition
Contact Us
Benjamin Miller Lab
MC 6-6141A
601 Elmwood Ave
Rochester, NY 14642