Structural Dynamics of Translation
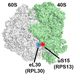
X-ray crystal structure of
bacterial 70S ribosome
The Ermolenko laboratory studies mechanisms of protein synthesis. The ribosome is a central component of cell metabolism, an extremely complex and highly dynamic machine. Although basic principles of protein synthesis have been established, many critical mechanisms that control translation remain poorly understood. We investigate the modulation of ribosome structural dynamics by translation factors and antibiotics, programmed ribosome frameshifting, and the regulation of translation by mRNA structure. In our studies, we combine biochemical approaches, single-molecule Förster Resonance Energy Transfer (smFRET) microscopy, and genetics. Our research on protein synthesis in bacterial, yeast, and human cells spans the spectrum of science from in silico to in vivo.

Dmitri N. Ermolenko, Ph.D.
Principal Investigator
- An smFRET assay to probe the impact of antibiotics on intersubunit rotation in eukaryotic ribosomes.; Methods in enzymology; Vol 725, pp. 77-103. 2025 Nov 11.
- 40S ribosomal subunits scan mRNA for the start codon by one-dimensional diffusion.; RNA (New York, N.Y.). 2025 Jul 31.
- Observing intersubunit dynamics in single yeast ribosomes.; bioRxiv : the preprint server for biology. 2025 Jun 08.
- 40S ribosomal subunits scan mRNA for the start codon by one-dimensional diffusion.; bioRxiv : the preprint server for biology. 2025 Jan 04.
News
Affiliations
- Biochemistry & Biophysics
- Biology
- Center for AIDS Research
- Center for RNA Biology: From Genome to Therapeutics
- NIH T32 Training Grant in Cellular, Biochemical & Molecular Sciences
- Biochemistry & Molecular Biology Ph.D. Program
- Biomedical Genetics and Genomics Ph.D. Program
- Biophysics, Structural & Computational Biology Ph.D. Program
June 12, 2024
Dmitri Ermolenko, PhD promoted to Professor
May 19, 2023
A Search Engine for mRNA: Algorithm Identifies Optimal Sequences to Improve COVID Vaccines
February 21, 2022
Congratulations Dr. Bao
February 21, 2022
Nature Communications Paper Published
Lab Photos
Contact Us
Ermolenko Lab
MC 3-8560
601 Elmwood Ave
Rochester, NY 14642
(585) 271-2683