Discovering new roles of mRNA secondary structure in eukaryotic translational control
Discovering new roles of mRNA secondary structure in eukaryotic translational control
We investigate how mRNA secondary structure regulates translation initiation in both eukaryotes (https://pmc.ncbi.nlm.nih.gov/articles/PMC6193969/; https://pmc.ncbi.nlm.nih.gov/articles/PMC9750134/; https://pmc.ncbi.nlm.nih.gov/articles/PMC12415969/) and bacteria (https://pmc.ncbi.nlm.nih.gov/articles/PMC6693549/). In collaboration with David Mathews’ laboratory (https://rna.urmc.rochester.edu/), we study the biological roles of conserved elements of secondary structure in the 5′ unstructured regions (5′ UTRs) of mRNA. We also investigate the interactions between the 5′ and 3′ mRNA ends in eukaryotic translational control. We have demonstrated that most mRNAs and long non-coding RNAs intrinsically fold, in the absence of proteins, into compact conformations characterized by short end-to-end distances, largely independent of RNA length (https://pmc.ncbi.nlm.nih.gov/articles/PMC6193969/). Multiple protein complexes that bridge the 5′ and 3′ ends of mRNA likely evolved to utilize this inherent RNA compactness (https://pmc.ncbi.nlm.nih.gov/articles/PMC8107001/). Beyond their mechanistic implications, our studies establish a quantitative framework for measuring, modeling, and engineering RNA end-to-end distances and secondary structure, with broad applications in basic research and biotechnology.
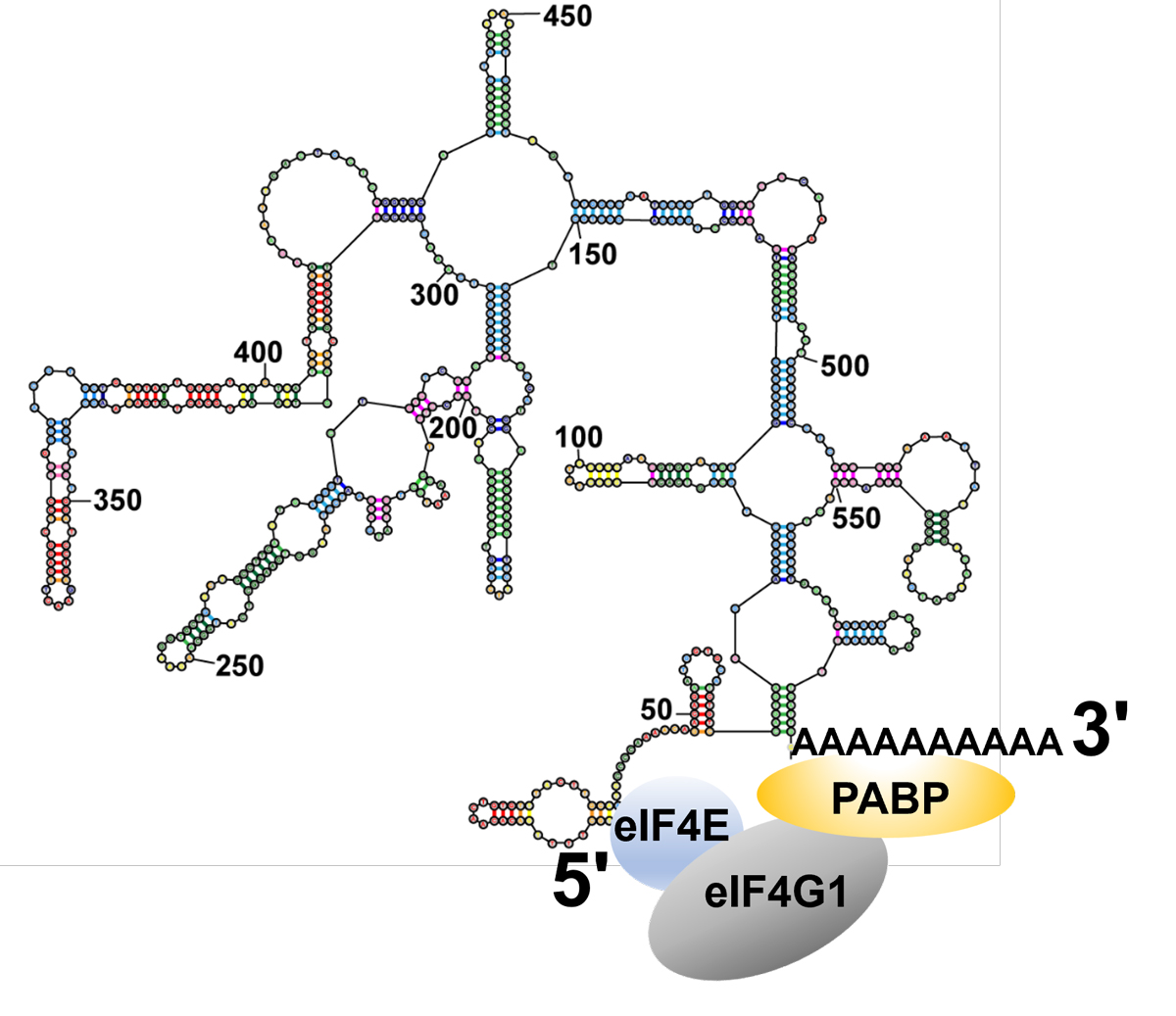
Figure legend: The intrinsic proximity of mRNA ends facilitates the formation of protein complexes that regulate translation initiation by bridging the two ends of mRNAs.