Studying structural rearrangements of the ribosome and translation factors
Studying structural rearrangements of the ribosome and translation factors
Protein synthesis is a multi-step process that is accompanied by intricate, large-scale structural rearrangements of the ribosome. To follow ribosome dynamics, we employ single-molecule Förster resonance energy transfer (smFRET) measurements in translation systems reconstituted from purified components. Using this approach, we observed in solution the rotational movement between the small and large ribosomal subunits of the bacterial ribosome, which was originally inferred from static cryo-EM structures. We have demonstrated that multiple translation factors and antibiotics modulate ribosome dynamics by selectively stabilizing either the rotated or nonrotated conformational states. More recently, we have developed a new smFRET assay to follow intersubunit rotation in eukaryotic (yeast) ribosomes (https://pmc.ncbi.nlm.nih.gov/articles/PMC12758802/). We employ our smFRET assays in single-molecule studies of pausing, stalling, and frameshifting during protein synthesis.
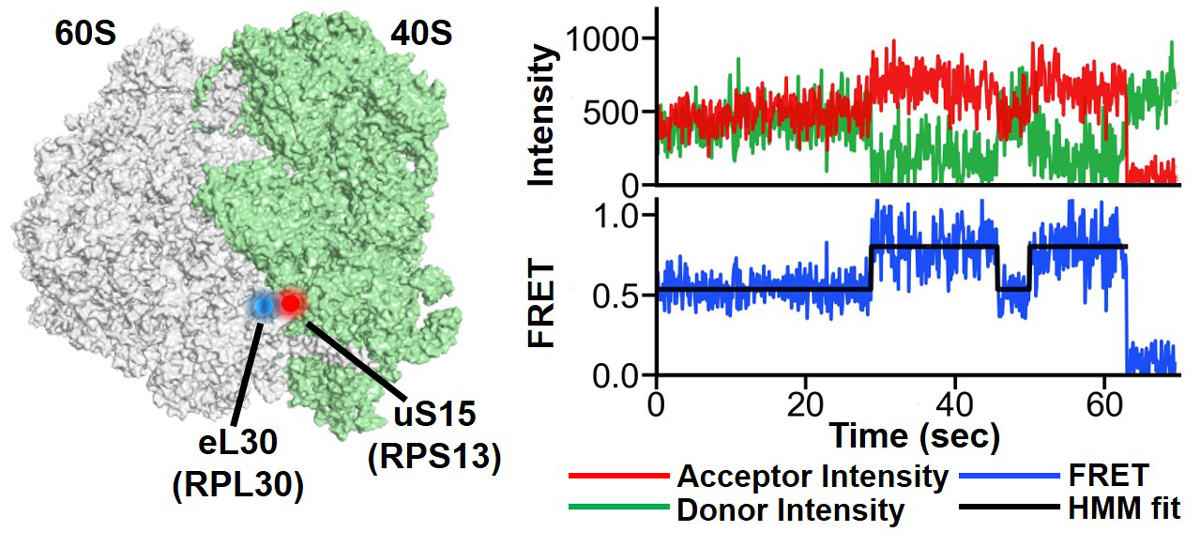
Figure legend: smFRET measurements of intersubunit rotation in yeast ribosomes