Carbon Nanotubes
Carbon Nanotubes
To facilitate high efficiency gene transfer into cells, arrays of carbon nanotubes are manufactured by template-based chemical vapor deposition (CVD), using commercially available anodized aluminum oxide membranes as template. These membranes contain pores, which determine the diameter and spacing of the carbon nanotubes. Carbon is deposited on all the surfaces of the membrane including the inner lumen of the pores, and after partial removal of the surface carbon and sacrificial membrane, the CNT tips are revealed. For our initial experiments we chose a Whatman AAO membrane with a pore diameter of 250 nm and a spacing of 200 nm between pores. The thickness of nanotube walls was controlled by CVD time, temperature and gas flow rate. After removing the excess carbon deposited on the membrane top surface with oxygen plasma, one side of the AAO membrane was partially etched with reactive ion etching (RIE) using boron trichloride to expose the tips of CNTs embedded in the membrane. The exposed CNT length was controlled by RIE etching time and RF power.
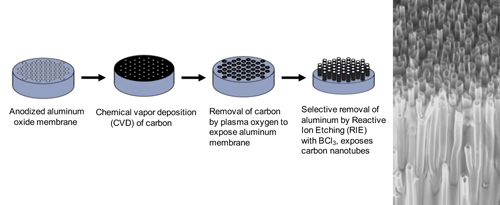
Manufacture of carbon nanotube arrays. Right image is scanning electron micrograph of
fractured edge of CNT device showing carbon tubes that extend through the plane of the array.
Our carbon nanotube array consists of thousands of closely-spaced, short carbon nanotubes, which provides an optimal surface for cell growth. Importantly, the basal layer of cells appears to engulf the nanotubes, which we hypothesize supports high efficiency gene transfer. Our first generation carbon nanotube arrays enable high efficiency gene transfer into immortalized cell lines, achieving 85% efficiency into HEK293 cells.
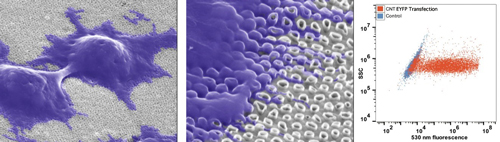

High efficiency gene transfer into HEK293 cells using carbon nanotube arrays. Cells grow
on carbon nanotube surface, engulfing carbon nanotubes (scanning electron microscopy
images). To quantify gene transfer, HEK293 cells were plated on the carbon nanotube
array, grown for 48 hr, and then transfected for 2 hr with plasmid encoding EYFP. Cells
were then cultured for 48 hr to allow for EYFP expression, and then harvested for analysis
by flow cytometry. Fluorescence is depicted in the X-axis, and cell number in Y-axis.
Current experiments are underway to develop second generation carbon nanotube arrays for gene transfer into cells that are not easily transfected by standard methods, including stem cells, iPSCs, and primary T-cells. Additionally, we are working to increase the size and complexity of cargo that can be transferred through carbon nanotube arrays, including mixed cargo of protein and nucleic acid.
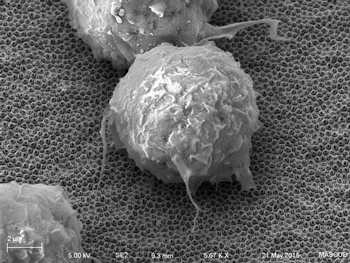
Scanning electron micrograph of macrophage
cell growing on carbon nanotube array.
These improved carbon nanotube arrays will be used for gene transfer into stem cells for enhanced CRISPR/Cas9 gene editing, for de-differentiating primary cells into iPSCs, for differentiating iPSCs into model cell systems, and for transferring chimeric antigen receptors into primary T-cells.