Elucidating mechanisms of programmed ribosome frameshifting
Elucidating mechanisms of programmed ribosome frameshifting
The translating ribosome must unfold mRNA secondary structure to feed single-stranded mRNA through the narrow mRNA channel of the small ribosomal subunit. Although the translating ribosome is a highly efficient helicase, certain RNA stem-loop structures can stall translation, triggering the No-Go mRNA decay pathway, inducing programmed ribosome pauses, and stimulating programmed ribosome frameshifting (PRF) in both bacteria and eukaryotes. PRF regulates viral gene expression and expands the viral proteome of retro- and coronaviruses, such as HIV and SARS-CoV-2. We address a fundamental question: how mRNA secondary structure interacts with the ribosome to regulate translation rate. We have found that rather than high thermodynamic stability, specific length and structure enable regulatory mRNA stem-loops to stall translation by forming inhibitory interactions with the ribosome (https://pmc.ncbi.nlm.nih.gov/articles/PMC8861025/). We have also discovered a new mechanism of ribosome pausing by demonstrating that mRNA stem-loops can transiently bind to the A site of the ribosome and sterically hinder tRNA binding (https://pmc.ncbi.nlm.nih.gov/articles/PMC7282821/). Our work informs the design of potential antiviral therapeutics targeting ribosome frameshifting utilized by human retro- and coronaviruses.
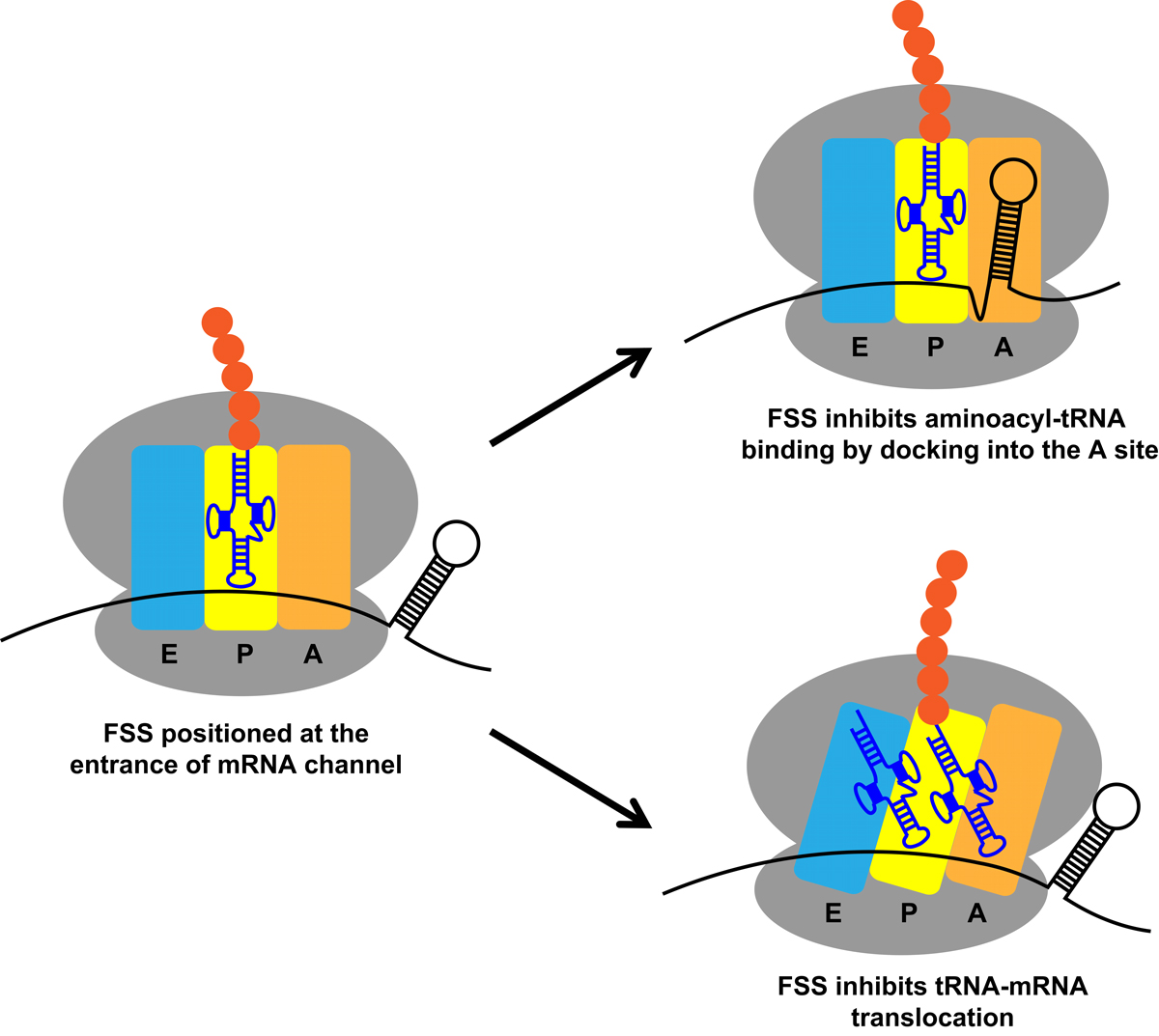
Figure legend: Mechanisms by which frameshift-inducing stem-loops or frameshift-stimulating sequences (FSSs) pause the ribosome during programmed ribosome frameshifting