Tolerance to drugs from echinocandin class: caspofungin, anidulafungin and micafungin
The loss and gain of one homolog
of a chromosome represents a novel
form of gene regulation.
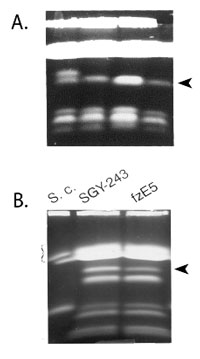
Our studies of the control of resistance to sorbose have provided a foundation on which to understand how this organism responds to antifungal drugs. When we generated mutants on medium supplemented with fluconazole, a major anticandidal drug from azole class of drugs, we found that these mutants are adapted to fluconazole due to alteration of copy number of a specific chromosome (Figure B). We also made the exciting discovery that Ch5 monosomy in mutants adapted to sorbose, alters, additionally, susceptibility to four major antifungal drugs, including increased tolerance to caspofungin that kills C. albicans in a fashion similar to sorbose. We, thus, generated caspofungin-tolerant mutants by a direct selection on medium supplemented with caspofungin and found several distinctive mechanisms of tolerance including:
- Ch5 monosomy similar to the sorbose-generated mutants
- A combination of one normal Ch5 and one iso-Ch5 with two right arms
- An aneuploidy-independent downregulation of a significant number of Ch5 genes on normal disomic Ch5s; these genes being the same as downregulated genes on the monosomic Ch5
We initially identified a total of three Ch5 genes PGA4, CHT2, and CSU51 encoding negative regulators of susceptibility to echinocandin drugs caspofungin and anidulafungin. However, the total number of such regulatory genes on Ch5 is yet to be established. Recently, we discovered five upregulated genes on Ch2 (positive regulators) and at least ten downregulated genes on Ch5 (negative regulators) that all change simultaneously in echinocandin-adapted mutants. These genes act to control surface exposure of the immunogenic cell wall epitope β-glucan, as well as to diminish its amount in the cell wall.
Caspofungin
In summary, we developed a new area of C. albicans research by discovering chromosomal alterations and linking aneuploidy, epigenetic regulation and responses to drugs and other environmental stresses. Our studies of laboratory resistance to toxic sorbose, fluconazole, and echinocandin drugs have provided a foundation on which to understand how C. albicans adapts to antifungals. Currently, the occurrence of various C. albicans chromosome alterations, including Ch5, is well documented in animal models and human patients. We are in the process of identifying many genes in C. albicans genome and in particular residing on Ch5 that contribute to evolution of clinical resistance to echinocandins.