Current Research Projects
Adaptable modularity in cellular functions
A./ Bacterial chemotaxis
Long, Z., Quaife B, Salman, H., Oltvai, Z.N. (2017) Cell-cell communication enhances bacterial chemotaxis toward external attractants. Sci. Rep., 7:12855; PMID: 28993669
- highlighted in Curr. Opin. Microbiol.: PMID: 29453124
Rashid, S., Long, Z., Singh, S., Kohram, M., Vashistha, H., Navlakha, S., Salman, H., Oltvai, Z.N, Bar-Joseph, Z., (2019) Adjustment in tumbling rates improves bacterial chemotaxis on obstacle-laden terrains. Proc. Natl. Acad. Sci. U.S.A., 116, 11770-11775; PMID: 31127043
- highlighted in Nat. Rev. Microbiol.: PMID: 31182826
B./ Cell metabolism
Singh, M., Warita, K., Warita, T., Faeder, J.R., Lee, R.E.C., Sant, S., Oltvai, Z.N. (2018) Shift from stochastic to spatially-ordered expression of serine-glycine synthesis enzymes in 3D microtumors. Sci. Rep., 8: 9388; PMID: 29925909 synopsys
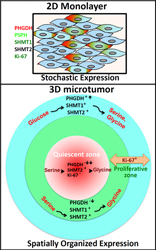 Network biology-based modeling of cell metabolism is constrained by the available genomic and functional -omic data that have almost exclusively been collected using 2-dimensional (2D) monolayer or liquid cell cultures. Yet, in reality, most cells exist as components of ecosystems in which they are responding dynamically to a cacophony of temporally changing signals from their environment and from other cells. In this paper, we show that monoclonal tumor cell 2D monolayers display a stochastic expression of PHGDH, the rate limiting enzyme of the serine-glycine synthesis pathway, that shift to a spatially-restricted expression pattern when the same monoclonal tumor cells are grown as 3D-microtumors. A mathematical model indicates that this expression pattern is an emergent property of this simple ecosystem, likely driven by intercellular paracrine signals from the hypoxic and relatively nutrient-deprived microtumor core. This paper clearly illustrates that studying cells in a microenvironment that recapitulates their natural ecosystem state is an essential requirement for future research on cell metabolism.
Network biology-based modeling of cell metabolism is constrained by the available genomic and functional -omic data that have almost exclusively been collected using 2-dimensional (2D) monolayer or liquid cell cultures. Yet, in reality, most cells exist as components of ecosystems in which they are responding dynamically to a cacophony of temporally changing signals from their environment and from other cells. In this paper, we show that monoclonal tumor cell 2D monolayers display a stochastic expression of PHGDH, the rate limiting enzyme of the serine-glycine synthesis pathway, that shift to a spatially-restricted expression pattern when the same monoclonal tumor cells are grown as 3D-microtumors. A mathematical model indicates that this expression pattern is an emergent property of this simple ecosystem, likely driven by intercellular paracrine signals from the hypoxic and relatively nutrient-deprived microtumor core. This paper clearly illustrates that studying cells in a microenvironment that recapitulates their natural ecosystem state is an essential requirement for future research on cell metabolism.
Delaying metastasis formation by synergistic inhibition of metabolic pathways
Warita, K., Warita, T., Beckwitt, C.H, Schurdak, M.E., Vazquez, A., Wells, A., Oltvai, Z.N., (2014) Statin-induced mevalonate pathway inhibition attenuates the growth of mesenchymal-like cancer cells that lack functional E-cadherin mediated cell cohesion. Sci. Rep., 4:7593, PMID: 25534349
Raghu, V., Beckwitt, C.H, Warita, K., Wells, A., Benos, P.V., Oltvai, Z.N., (2018) Biomarker identification for statin sensitivity of cancer cell lines. Biochem. Biophys. Res. Comm., 495: 659-665; PMID: 29146185
Beckwitt, C.H., Clark, A.M., Ma, B., Whaley, D., Oltvai, Z.N., Wells, A. (2018) Statins attenuate outgrowth of breast cancer metastases. Br. J. Cancer, 119, 1094–1105; PMID: 30401978
Ishikawa, T., Hosaka, Y.Z., Beckwitt, C.H., Wells, A., Oltvai, Z.N., Warita, K. (2018) Concomitant attenuation of HMG-CoA reductase expression potentiates the cancer cell growth-inhibitory effect of statins and expands their efficacy in tumor cells with epithelial characteristics. Oncotarget, 9: 29304-29315; PMID: 30034619
Warita, K., Ishikawa, T., Sugiura, A., Tashiro, J., Shimakura, H., Hosaka, Y.Z., Ohta, K., Warita, T. and Oltvai, Z.N. (2021) Concomitant attenuation of HMGCR expression and activity enhances the growth inhibitory effect of atorvastatin on TGF-β-treated epithelial cancer cells. Sci. Rep., 11: 12763; PMID: 34140545
Methods for pathogenicity and targeted therapy resistance assessment of genomic variants
Ponzoni, L., Penaherrera, D.A., Oltvai, Z.N., Bahar, I. (2020) Rhapsody: Predicting the pathogenicity of human missense variants. Bioinformatics, 36, 3084-3092; PMID: 32101277
Oltvai, Z.N., Smith, E.A., Wiens, K., Nogee, L.M., Luquette, M., Nelson, A.C., Wikenheiser-Brokamp, K.A. (2020) Neonatal respiratory failure due to novel compound heterozygous mutations in the ABCA3 lipid transporter. Cold Spring Harb. Mol. Case Stud., 6:a005074; PMID: 32532878
- Highlighted on the cover: http://molecularcasestudies.cshlp.org/content/6/3.cover-expansion
Oltvai Z.N., Harley, S.E., Koes, D., Michel, S., Warlick, E.D., Nelson, A.C., Yohe, S. and Mroz, P (2021). Assessing acquired resistance to IDH1 inhibitor therapy by full exon IDH1 sequencing and structural modeling, Cold Spring Harb. Mol. Case Stud., 7:a006007, PMID: 33832922
- Highlighted on the cover: http://molecularcasestudies.cshlp.org/content/7/2.cover-expansion