Projects
The Biological Roles of tRNA Modifications
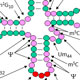 tRNAs are extensively modified in all organisms; in the yeast Saccharomyces cerevisiae, for example, cytoplasmic tRNAs have 25 biochemically distinct modifications, and the average tRNA has 13 modified residues. Modifications are highly conserved in eukaryotes, including humans, and their biological roles are a major topic of study. Modifications in and around the anticodon loop generally affect aspects of translation, whereas modifications in the main body of the tRNA often affect tRNA folding or stability. However, the precise roles of many modifications remain poorly understood, and some of these are under investigation in this lab.
tRNAs are extensively modified in all organisms; in the yeast Saccharomyces cerevisiae, for example, cytoplasmic tRNAs have 25 biochemically distinct modifications, and the average tRNA has 13 modified residues. Modifications are highly conserved in eukaryotes, including humans, and their biological roles are a major topic of study. Modifications in and around the anticodon loop generally affect aspects of translation, whereas modifications in the main body of the tRNA often affect tRNA folding or stability. However, the precise roles of many modifications remain poorly understood, and some of these are under investigation in this lab.
Learn more about The Biological Roles of tRNA Modifications
Investigation of the Specificity of Modification Enzymes
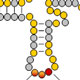 A second major problem of interest to us is the nature of the substrate specificity of modification enzymes. Since tRNA modifications are critical for the proper folding and stability of tRNA, and for high fidelity decoding of mRNA during translation, mechanisms to ensure accurate modification of each tRNA species must exist. This specificity can be crucial because failure of modification frequently leads to nonfunctional or poorly functioning tRNA, and because inadvertent modification of the incorrect tRNA species can have deleterious effects on the cell. Modification enzymes are highly specific for certain tRNA species and for specific residues within these tRNAs, and identifying the sequence and/or structural determinants for modification is important for understanding and possibly manipulating the biology.
A second major problem of interest to us is the nature of the substrate specificity of modification enzymes. Since tRNA modifications are critical for the proper folding and stability of tRNA, and for high fidelity decoding of mRNA during translation, mechanisms to ensure accurate modification of each tRNA species must exist. This specificity can be crucial because failure of modification frequently leads to nonfunctional or poorly functioning tRNA, and because inadvertent modification of the incorrect tRNA species can have deleterious effects on the cell. Modification enzymes are highly specific for certain tRNA species and for specific residues within these tRNAs, and identifying the sequence and/or structural determinants for modification is important for understanding and possibly manipulating the biology.
Learn more about Investigation of the Specificity of Modification Enzymes