URMC / Labs / Phizicky Lab / Projects / Investigation of the Specificity of Modification Enzymes
Investigation of the Specificity of Modification Enzymes
We previously examined the specificity for G-1 addition to tRNAHis and the specificity of tRNA variants for RTD, and recently we have focused on the specificity of modification enzymes for C32. We are interested in the C32 modification because this residue is in the anticodon loop and is implicated in translation, and because C32 can have three different fates in different tRNAs: it is modified to m3C in 6 tRNA species, it is modified to Cm in 3 tRNAs, and it is unmodified in 14 other tRNA species.
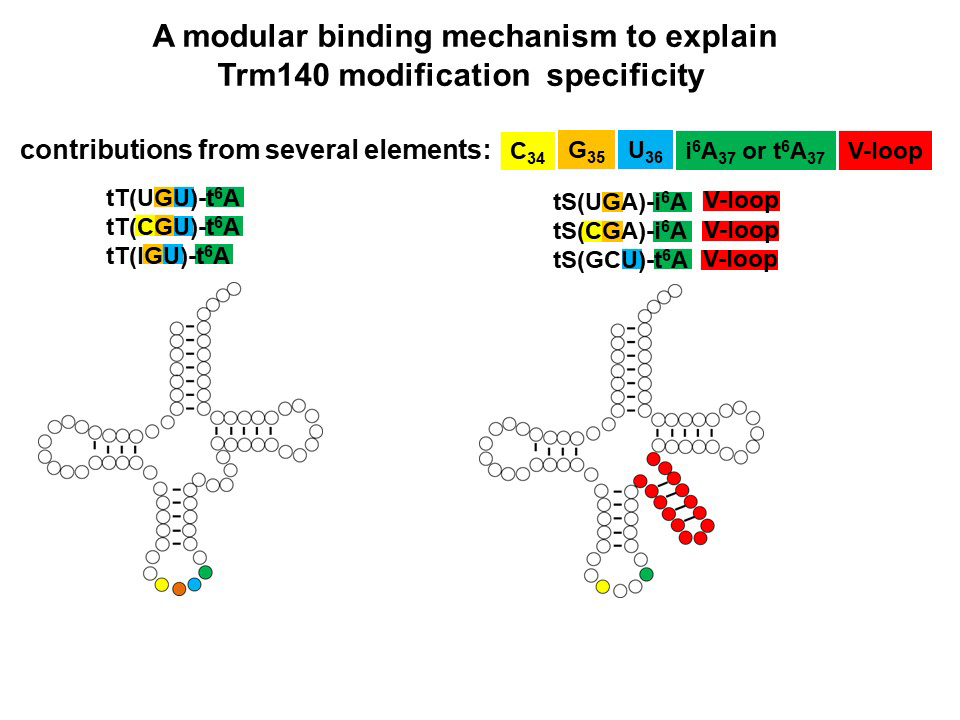
Our recent results show that the specificity of S. cerevisiae Trm140 for m3C modification is complex. Trm140 recognizes and modifies its three tRNAThr substrates (with anticodons IGU, CGU, and UGU) based entirely on recognition of the anticodon residues G35 and U36 and the adjacent t6A37 residue Mutation of any of any of these residues eliminates modification in vivo and binding in vitro, and introduction of these residues to another tRNA is sufficient for both binding and modification. Remarkably, however, Trm140 recognition of the three tRNASer substrates (with anticodons UGA, CGA, and GCU) is completely different. In this case, m3C modification requires seryl-tRNA synthetase and the distinctive variable loop of tRNASer in vitro and in vivo, and is stimulated by t6A37 or i6A37 as well as by C34 and G35. These seemingly very different modes of Trm140 recognition of tRNAThr and tRNASer substrates appear to be widely conserved in Ascomycota, but Trm140 has duplicated into two or more paralogs in other fungi and in metazoans for separate modification of the two tRNA families.
Current work is focused on understanding the tRNA determinants that are recognized by Trm7 and its partner proteins, which also recognize C32 of its substrate tRNAs, as well as the nearby anticodon residue N34, and also have no obvious recognition elements in common.
« back to all projects