Collaborative Research
Define the identity and properties of inhibitory codon combinations

Although accurate and efficient translation of mRNA into proteins is crucially important for the cell, not all of the 61 codons that specify insertion of amino acids into nascent polypeptides behave equally in this process. In fact, almost all synonymous codons, i.e. those that specify the same amino acid, are used at very different frequencies within most organisms, including the yeast S. cerevisiae.
High Throughput Purification of Proteins for Biochemical and Structural Analysis

Recent work in this lab and the lab of E. Grayhack has focused on establishing yeast as the eukaryote of choice for high throughput purification of yeast and other eukaryotic proteins for biochemical analysis and structural biology. Although expression in E. coli has been used for biochemical analysis and for the determination of a large number of protein structures, expression of eukaryotic proteins in E. coli often results in limited solubility, as well as in the absence of post-translational modifications. To establish yeast as an organism for high throughput cloning, expression and purification, we have developed several methods.
Post-transcriptional Gene Regulation by Double-stranded RNA-binding Proteins
We have found that STAU1, which is a double-stranded RNA binding protein, recruits the NMD factor UPF1 to certain mRNA 3'-untranslated regions (3'UTRs) so as to elicit SMD in a translation-dependent mechanism (reviewed in Park and Maquat, 2013, Wiley Interdiscip Rev RNA 4:423-435). Using microarray analyses, we have identified a number of mRNAs that are naturally down-regulated by SMD.
Regulation of Protein Synthesis by mRNA Structure

The central dogma of Biology is that DNA is used to make messenger RNA (mRNA), which is used to make proteins. Over the last decade, multiple findings have illuminated the importance of the regulation of protein expression at the level of mRNA translation. mRNA is no longer considered a simple courier of genetic information between DNA and protein. RNA can fold into an extensive secondary and, in many instances, tertiary structure. In recent years, numerous studies began to reveal that structured elements within the mRNA play a critical role in modulating the flow of genetic information from DNA to protein.
The Biological Roles of tRNA Modifications
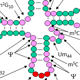
tRNA is the most highly modified class of RNA species, and modifications are found in tRNAs from all organisms that have been examined. Yeast cytoplasmic tRNAs have 25 biochemically distinct modifications, and the average tRNA has 13 modified residues. These modifications are highly conserved in eukaryotes, including humans, and many of the corresponding genes have been identified in this and other labs. It is known from previous work in the field that several modifications in and around the anticodon loop have crucial roles in various aspects of translation, while several other modifications remote from the anticodon loop have specific roles in tRNA folding and/or stability. However, the precise roles of many other individual modifications are only poorly understood, and are under investigation in our lab.