Research Projects
Studying structural rearrangements of the ribosome and translation factors
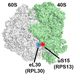 Protein synthesis is a multi-step process that is accompanied by intricate, large-scale structural rearrangements of the ribosome. To follow ribosome dynamics, we employ single-molecule Förster resonance energy transfer (smFRET) measurements in translation systems reconstituted from purified components. Using this approach, we observed in solution the rotational movement between the small and large ribosomal subunits of the bacterial ribosome, which was originally inferred from static cryo-EM structures. We have demonstrated that multiple translation factors and antibiotics modulate ribosome dynamics by selectively stabilizing either the rotated or nonrotated conformational states. More recently, we have developed a new smFRET assay to follow intersubunit rotation in eukaryotic (yeast) ribosomes (https://pmc.ncbi.nlm.nih.gov/articles/PMC12758802/). We employ our smFRET assays in single-molecule studies of pausing, stalling, and frameshifting during protein synthesis.
Protein synthesis is a multi-step process that is accompanied by intricate, large-scale structural rearrangements of the ribosome. To follow ribosome dynamics, we employ single-molecule Förster resonance energy transfer (smFRET) measurements in translation systems reconstituted from purified components. Using this approach, we observed in solution the rotational movement between the small and large ribosomal subunits of the bacterial ribosome, which was originally inferred from static cryo-EM structures. We have demonstrated that multiple translation factors and antibiotics modulate ribosome dynamics by selectively stabilizing either the rotated or nonrotated conformational states. More recently, we have developed a new smFRET assay to follow intersubunit rotation in eukaryotic (yeast) ribosomes (https://pmc.ncbi.nlm.nih.gov/articles/PMC12758802/). We employ our smFRET assays in single-molecule studies of pausing, stalling, and frameshifting during protein synthesis.
Learn more about Studying structural rearrangements of the ribosome and translation factors
Elucidating mechanisms of programmed ribosome frameshifting

The translating ribosome must unfold mRNA secondary structure to feed single-stranded mRNA through the narrow mRNA channel of the small ribosomal subunit. Although the translating ribosome is a very efficient helicase, certain RNA stem-loop structures can stall translation to trigger the No-Go mRNA decay pathway, induce programmed ribosome pauses, and stimulate programmed ribosome frameshifting (PRF) in both bacteria and eukaryotes. PRF regulates viral gene expression and expands the viral proteome of retro- and coronaviruses, such as HIV and SARS-CoV-2. We address a fundamental question: how mRNA secondary structure interacts with the ribosome to regulate the rate of translation. We have found that rather than high thermodynamic stability, specific length and structure enable regulatory mRNA stem-loops to stall translation by forming inhibitory interactions with the ribosome (https://pmc.ncbi.nlm.nih.gov/articles/PMC8861025/). We have also discovered a new mechanism of ribosome pausing by demonstrating that mRNA stem-loops can transiently bind to the A site of the ribosome and sterically hinder tRNA binding (https://pmc.ncbi.nlm.nih.gov/articles/PMC7282821/). Our work informs the design of potential antiviral therapeutics targeting ribosome frameshifting utilized by human retro- and coronaviruses.
Learn more about Elucidating mechanisms of programmed ribosome frameshifting
Discovering new roles of mRNA secondary structure in eukaryotic translational control
 We investigate how mRNA secondary structure regulates translation initiation in both eukaryotes (https://pmc.ncbi.nlm.nih.gov/articles/PMC6193969/; https://pmc.ncbi.nlm.nih.gov/articles/PMC9750134/; https://pmc.ncbi.nlm.nih.gov/articles/PMC12415969/) and bacteria (https://pmc.ncbi.nlm.nih.gov/articles/PMC6693549/). In collaboration with David Mathews’ laboratory (https://rna.urmc.rochester.edu/), we study the biological roles of conserved elements of secondary structure in the 5′ unstructured regions (5′ UTRs) of mRNA. We also investigate the interactions between the 5′ and 3′ mRNA ends in eukaryotic translational control. We have demonstrated that most mRNAs and long non-coding RNAs intrinsically fold, in the absence of proteins, into compact conformations characterized by short end-to-end distances, largely independent of RNA length (https://pmc.ncbi.nlm.nih.gov/articles/PMC6193969/). Multiple protein complexes that bridge the 5′ and 3′ ends of mRNA likely evolved to utilize this inherent RNA compactness (https://pmc.ncbi.nlm.nih.gov/articles/PMC8107001/). Beyond their mechanistic implications, our studies establish a quantitative framework for measuring, modeling, and engineering RNA end-to-end distances and secondary structure, with broad applications in basic research and biotechnology.
We investigate how mRNA secondary structure regulates translation initiation in both eukaryotes (https://pmc.ncbi.nlm.nih.gov/articles/PMC6193969/; https://pmc.ncbi.nlm.nih.gov/articles/PMC9750134/; https://pmc.ncbi.nlm.nih.gov/articles/PMC12415969/) and bacteria (https://pmc.ncbi.nlm.nih.gov/articles/PMC6693549/). In collaboration with David Mathews’ laboratory (https://rna.urmc.rochester.edu/), we study the biological roles of conserved elements of secondary structure in the 5′ unstructured regions (5′ UTRs) of mRNA. We also investigate the interactions between the 5′ and 3′ mRNA ends in eukaryotic translational control. We have demonstrated that most mRNAs and long non-coding RNAs intrinsically fold, in the absence of proteins, into compact conformations characterized by short end-to-end distances, largely independent of RNA length (https://pmc.ncbi.nlm.nih.gov/articles/PMC6193969/). Multiple protein complexes that bridge the 5′ and 3′ ends of mRNA likely evolved to utilize this inherent RNA compactness (https://pmc.ncbi.nlm.nih.gov/articles/PMC8107001/). Beyond their mechanistic implications, our studies establish a quantitative framework for measuring, modeling, and engineering RNA end-to-end distances and secondary structure, with broad applications in basic research and biotechnology.