Early Life Exposures and Epigenetic Regulation
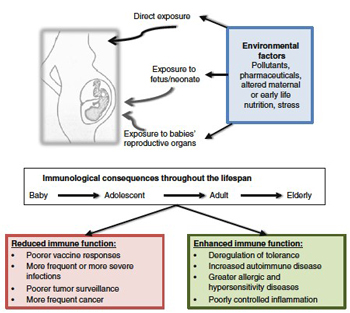
Exposure to environmental pollutants early in life is an often overlooked but important contributor to changes in immune system functions later in life.
Using clinically relevant mouse models and environmentally relevant exposures, we have several on-going projects to define cellular effects of early life exposure that causes long-lasting alterations to the function of the immune system. Exposures include small molecules that bind to the AHR, such as dioxins, and endocrine disrupting chemicals.
For example, by integrating contemporary immunology and genome-wide approaches, we have shown that inappropriate AHR activation during development causes durable functional changes in immune cells that are, at least in part, due to epigenetic mechanisms, such as alterations in DNA methylation. Moreover, we have exciting evidence that using nutritional supplements and pharmacological agents that these perturbations can be modulated later in life.
In other studies, we are examining how exposure to environmental chemicals, such as polychlorinated biphenyls (PCBs) and polyfluoroalkyl substances (PFAS), affect human immune defenses during infancy, childhood and adolescence using an established human cohorts.