Projects
The Role of the Integrator Complex in Gene Expression Regulation
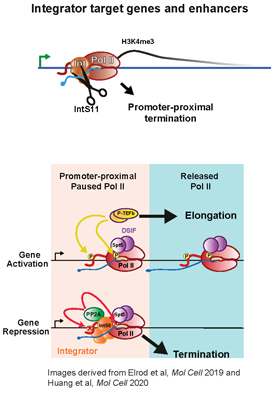 The Integrator Complex was discovered in 2005 as a 14 membered group of proteins that are required for the 3' end formation of small nuclear RNA. The Integrator Complex is thought to interact with machinery at the snRNA promoter including members of the SNAP complex, which leads Integrator association with the CTD of RNAPII. Once Integrator is loaded onto RNAPII, it will cleave the pre-snRNA using its catalytic RNA endonuclease subunit: IntS11. More recently, Integrator has been shown to function in the pause/release of RNAPII at many protein-coding genes. We and others have found that Integrator utilizes both its RNA endonuclease activity through IntS11 as well as an associated PP2A phosphatase to facilitate termination of promoter-proximal paused RNAPII. The Wagner laboratory is interested in determining the mechanism of Integrator action at these various gene types. To that end, we have developed model systems in both human and Drosophila cells in address its various unknowns. This project will enable a student to receive training in many aspects of biochemistry and molecular biology. Moreover, our group is highly collaborative enabling access to groups with expertise in both functional genomics and structural biology.
The Integrator Complex was discovered in 2005 as a 14 membered group of proteins that are required for the 3' end formation of small nuclear RNA. The Integrator Complex is thought to interact with machinery at the snRNA promoter including members of the SNAP complex, which leads Integrator association with the CTD of RNAPII. Once Integrator is loaded onto RNAPII, it will cleave the pre-snRNA using its catalytic RNA endonuclease subunit: IntS11. More recently, Integrator has been shown to function in the pause/release of RNAPII at many protein-coding genes. We and others have found that Integrator utilizes both its RNA endonuclease activity through IntS11 as well as an associated PP2A phosphatase to facilitate termination of promoter-proximal paused RNAPII. The Wagner laboratory is interested in determining the mechanism of Integrator action at these various gene types. To that end, we have developed model systems in both human and Drosophila cells in address its various unknowns. This project will enable a student to receive training in many aspects of biochemistry and molecular biology. Moreover, our group is highly collaborative enabling access to groups with expertise in both functional genomics and structural biology.
Alternative Polyadenylation in Cancer and other Human Disease Settings
 In non-proliferating or non-transformed cells, the position of cleavage and polyadenylation is located as distal as possible relative to the stop codon. The use of this distal poly(A) site (dPAS) ensures maximal negative regulation by microRNAs as well as AU-rich destabilizing elements. The translational output of these longer mRNAs is therefore under more exquisite control by the cell. However, during the processes of transformation, tumor progression, or enhanced proliferation; the decision of where to put the poly(A) tail is altered and a more proximal poly(A) site (pPAS) is chosen. This typically does not change sequences within the open reading frame, the reduced 3'UTR length removes negative regulatory elements increasing protein expression. In the Wagner laboratory, we are focused on the mechanism of this change and the critical target mRNAs that undergo this process. We use a combination of cellular and in vivo models to investigate APA as it relates to tumor biology.
In non-proliferating or non-transformed cells, the position of cleavage and polyadenylation is located as distal as possible relative to the stop codon. The use of this distal poly(A) site (dPAS) ensures maximal negative regulation by microRNAs as well as AU-rich destabilizing elements. The translational output of these longer mRNAs is therefore under more exquisite control by the cell. However, during the processes of transformation, tumor progression, or enhanced proliferation; the decision of where to put the poly(A) tail is altered and a more proximal poly(A) site (pPAS) is chosen. This typically does not change sequences within the open reading frame, the reduced 3'UTR length removes negative regulatory elements increasing protein expression. In the Wagner laboratory, we are focused on the mechanism of this change and the critical target mRNAs that undergo this process. We use a combination of cellular and in vivo models to investigate APA as it relates to tumor biology.
Use of PolyA-Click-seq (PAC-seq) to Investigate Disease Relevant Gene Expression and Alternative Polyadenylation
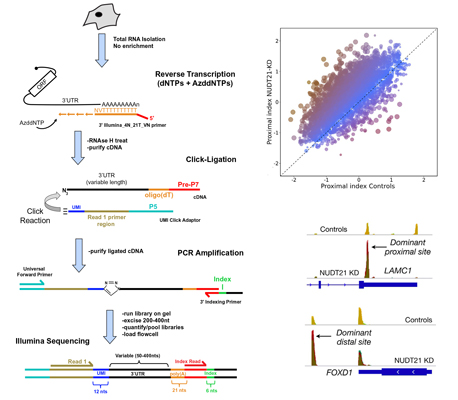 In collaboration with the Routh laboratory (UTMB-Galveston), we developed a novel sequencing technology called PolyA-Click-seq or PAC-seq. This library preparation has many advantages of current approaches in that it enriches for sequencing near the 3' ends of genes and is highly cost-effective. We use PAC-seq to monitor alternative polyadenylation events genome-wide as well as to measure differential gene expression. Students and postdoctoral fellows will have the opportunity to use PAC-seq in their own projects are as part of collaborations with other groups.
In collaboration with the Routh laboratory (UTMB-Galveston), we developed a novel sequencing technology called PolyA-Click-seq or PAC-seq. This library preparation has many advantages of current approaches in that it enriches for sequencing near the 3' ends of genes and is highly cost-effective. We use PAC-seq to monitor alternative polyadenylation events genome-wide as well as to measure differential gene expression. Students and postdoctoral fellows will have the opportunity to use PAC-seq in their own projects are as part of collaborations with other groups.