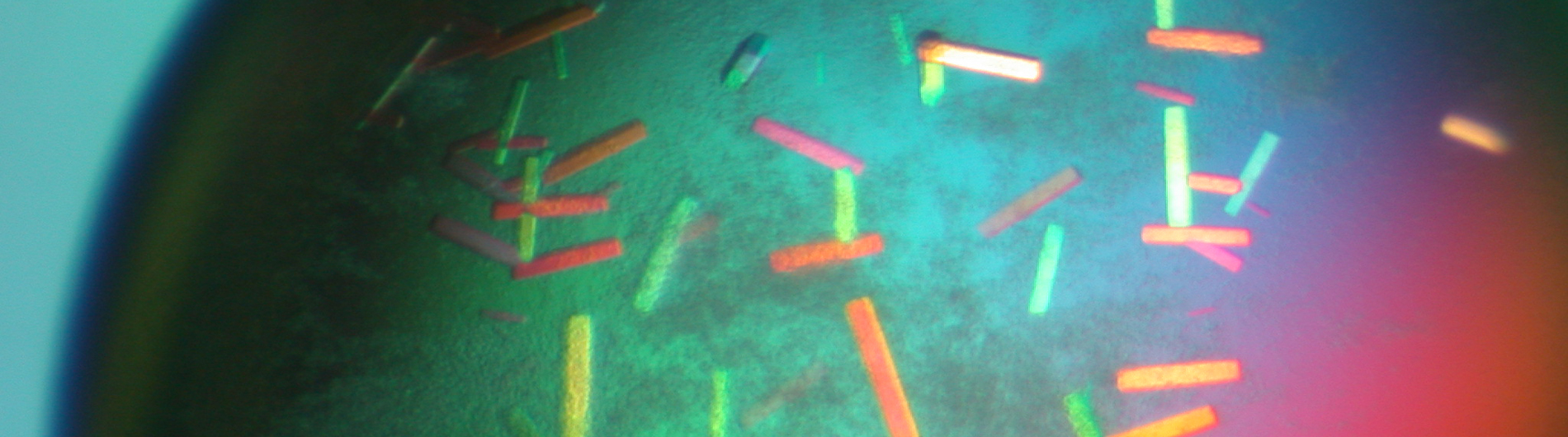
Structure and Function of Non-Coding RNAs in bacteria and HIV-1
Bacterial and viral infections pose grave threats to public health, and have reached epidemic proportions in some countries. Today’s physicians-in-training are likely to see chronic infections in their future medical practices, due to the emergence of drug resistance that continues to deplete the arsenal of antibiotics and antivirals once available for routine treatments. To revitalize this armamentarium, it is necessary to expand our knowledge of bacterial physiology and host defense with the goal of identifying novel therapeutic targets and strategies.
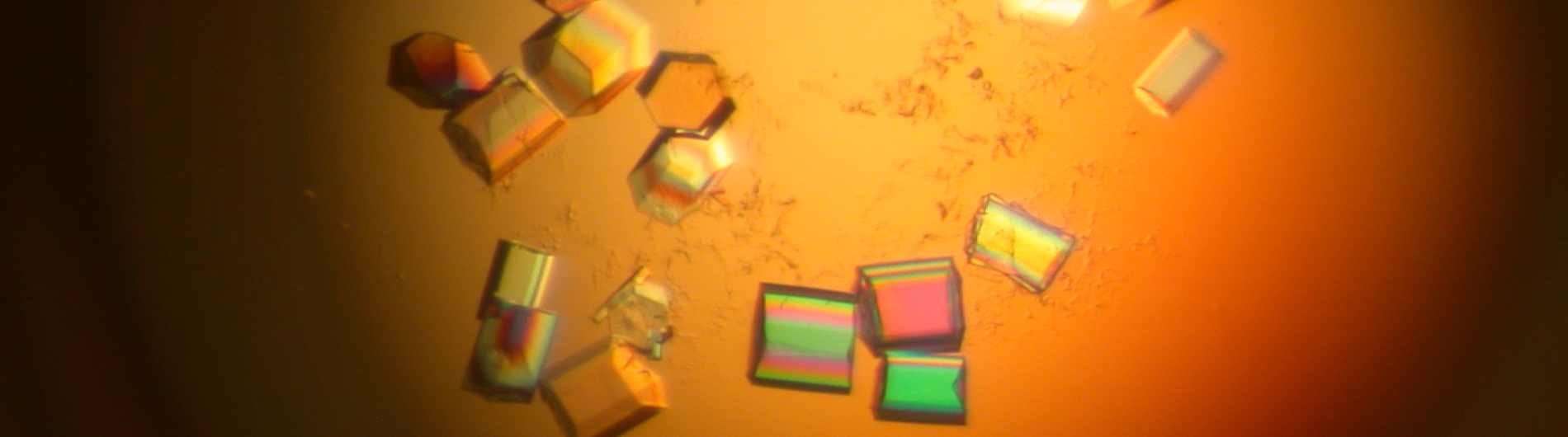
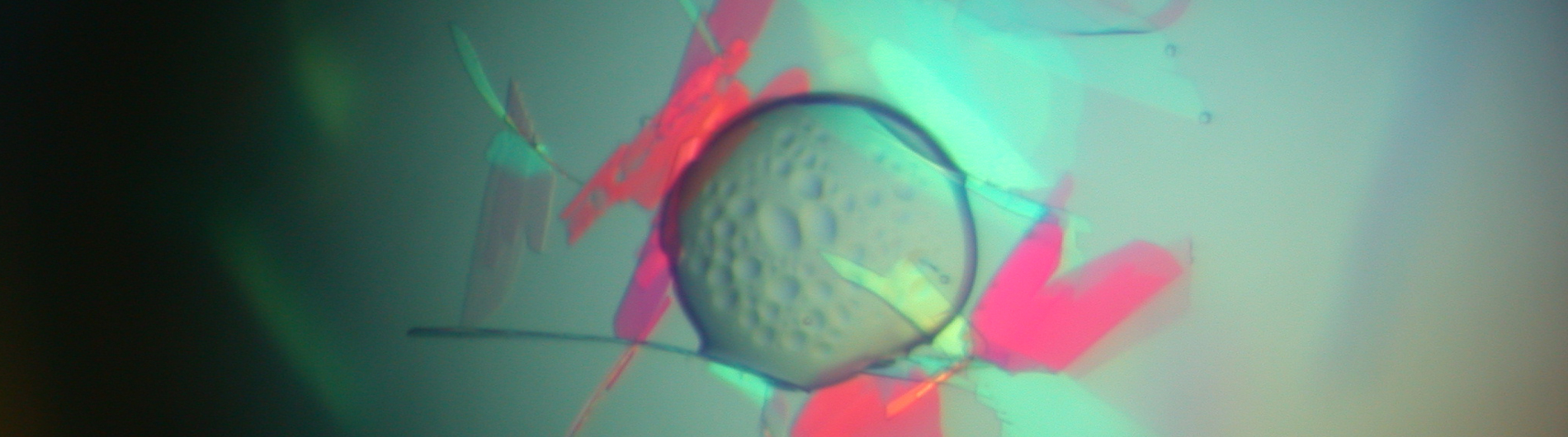
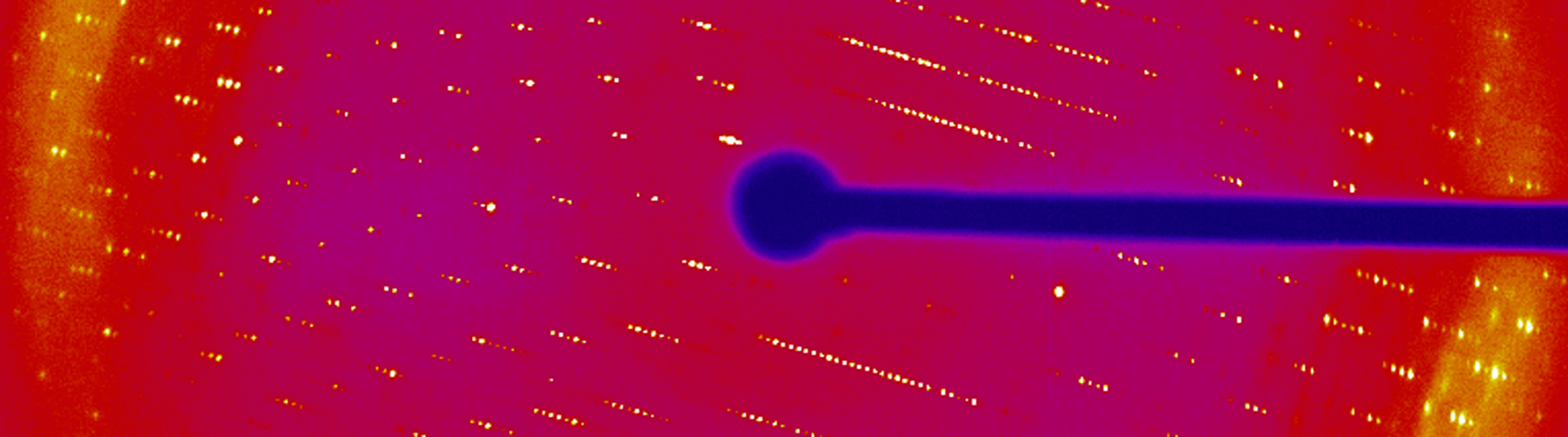
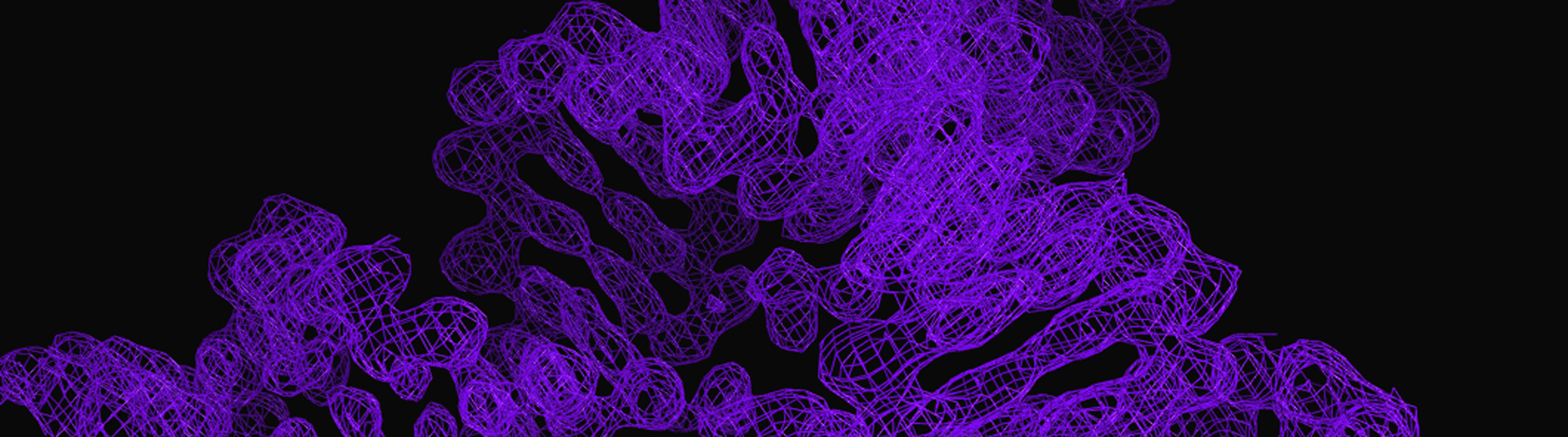
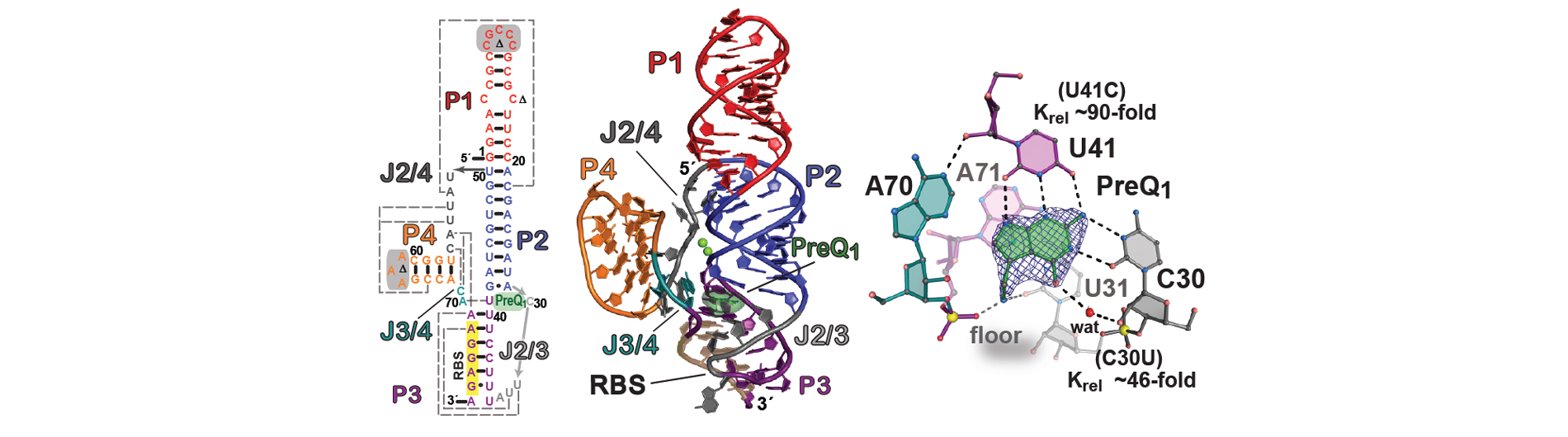
The Wedekind lab is working to understand RNA-mediated gene regulation by non-protein coding RNAs known as riboswitches, which hold promise as novel antibiotic targets. Although present in all domains of life, riboswitches are most prominent in bacteria where they can regulate as many as 5% of genes. An immediate goal of our research is to understand how these regulatory RNAs fold into complex three-dimensional structures that bind cognate small-molecule effectors to bring about gene “on” or gene “off” conformational states. Understanding riboswitch architecture is a first step toward the rational design of small molecules that can manipulate bacterial growth and virulence.
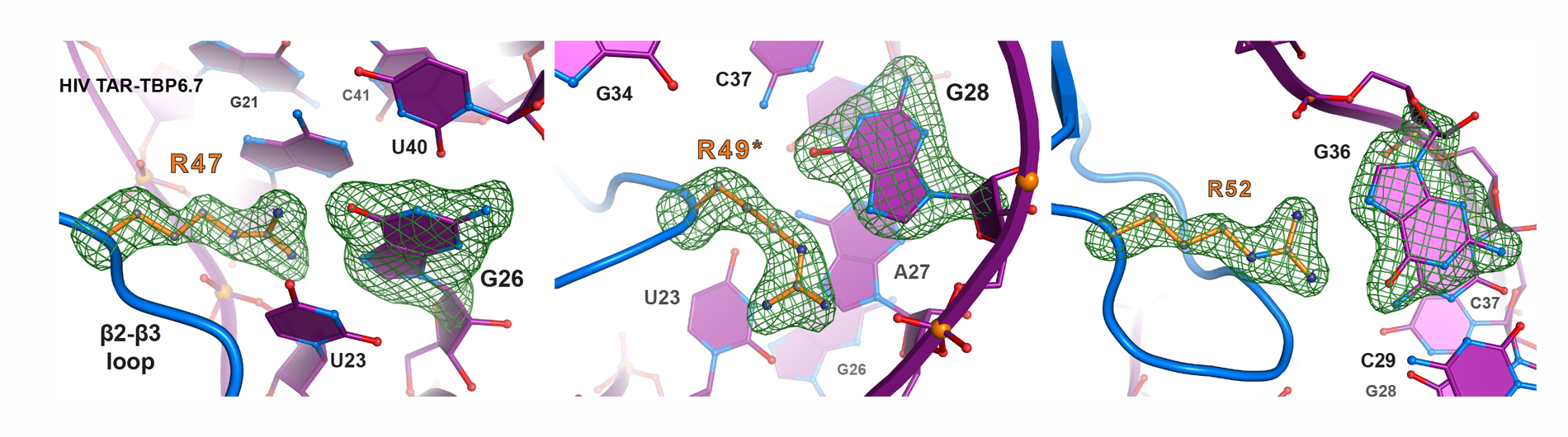
We are also investigating the use of lab-evolved proteins to control HIV-1 transcription by examining how variants of the U1A protein bind to the viral trans-activation response element (TAR) RNA. Our goal is to understand the basis of TAR RNA recognition by a peptide scaffold with the goal of exploiting this platform to develop a novel class of antiviral therapeutics.
Our lab utilizes a broad range of biophysical, biochemical, and computational approaches. We are also highly collaborative. We are currently funded by research grants from the National Institutes of Health/NIGMS (R01 GM063162 & R01 GM123864), and NIH/NCRR award S10 RR026501 for X-ray instrumentation housed in the Structural Biology and Biophysics Facility. We are grateful for two new NIH instrument supplements GM063162-14S1 ($63,916) and GM123864-02S1($129,175) that will be used to acquire a new ÄKTA Pure chromatography system (GE Healthcare) and cold cabinet, and a new low-volume PEAQ ITC (Malvern Panalytical). These items will be housed and maintained in the Wedekind Lab.

Joseph E. Wedekind, Ph.D.
Principal Investigator
Journal Covers







Publications
View All Publications- Knotty is nice: metabolite binding and RNA-mediated gene regulation by the preQ1 riboswitch family.; The Journal of biological chemistry. 2024 Oct 30.
- Cyclic peptides targeting the SARS-CoV-2 programmed ribosomal frameshifting RNA from a multiplexed phage display library.; Chemical science. 2024 Oct 17.
- Two riboswitch classes that share a common ligand-binding fold show major differences in the ability to accommodate mutations.; Nucleic acids research. 2024 Oct 16.
- Structure and function analysis of a type III preQ1-I riboswitch from E. coli reveals direct metabolite-sensing by the Shine-Dalgarno sequence.; The Journal of biological chemistry. 2023 Sep 01.
News
Affiliations
February 12, 2021
Wedekind lab research featured on the cover of JBC “The Year in JBC: 2020" issue
November 9, 2018
Wedekind Co-Organizes Nucleic Acids Session at American Crystallographic Association Annual Meeting in Toronto
January 26, 2016
Strong Star Certificate of Appreciation Awarded to Dr. Jermaine “JJ" Jenkins
Contact Us
Wedekind Lab
601 Elmwood Ave
Rochester, NY 14642