Research Projects
Human life depends on pre-mRNA splicing for cellular viability, differentiation and responses to changing physiology or environment. A major focus of my laboratory is to understand at a molecular level how the splicing machinery identifies sites for excision from gene transcript RNAs, which in turn changes the proteins produced. We have characterized the three-dimensional shapes of human splicing proteins recognizing one another and the gene transcript RNA at high resolution by X-ray crystallography complemented by molecular biology in human cells. Through this research, we identify a network of interactions responsible for recognizing human splice sites. The broader impact of this work for human disease is emphasized by the severe defects in pre-mRNA splicing that accompany most human hematologic malignancies and many metabolic disorders, as well as the dependence of HIV-1 and other complex retroviruses on RNA splicing for infectivity. Specific projects include:
Molecular Recognition During pre-mRNA Splicing
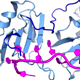 The overall goal of this research is to understand how essential ternary complexes of SF1, U2AF65, and U2AF35 splicing factors recognizes 3´ splice sites and initiates the process of spliceosome assembly.
The overall goal of this research is to understand how essential ternary complexes of SF1, U2AF65, and U2AF35 splicing factors recognizes 3´ splice sites and initiates the process of spliceosome assembly.
Learn more about Molecular Recognition During pre-mRNA Splicing
Molecular Actions of Prevalent U2AF1 Mutations in Myelodysplastic Syndromes
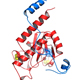 Specific mutations in the U2AF1 proto-oncogene, which encodes the U2AF35 protein, are prevalent among patients with hematological malignancies, including 10-12% of patients with myelodysplastic syndrome (MDS) without ring sideroblasts and 8-11% of patients with chronic myelomonocytic leukemia (CMML).
Specific mutations in the U2AF1 proto-oncogene, which encodes the U2AF35 protein, are prevalent among patients with hematological malignancies, including 10-12% of patients with myelodysplastic syndrome (MDS) without ring sideroblasts and 8-11% of patients with chronic myelomonocytic leukemia (CMML).
Learn more about Molecular Actions of Prevalent U2AF1 Mutations in Myelodysplastic Syndromes
Structural Control of Human Co-factors for Retroviral Gene Expression
 Viruses such as HIV-1 use the human machinery to produce its RNA transcripts for protein expression and ultimately genomic replication. How viruses hijack cellular processes through interactions with host macromolecules is a fundamental question in biology and medicine.
Viruses such as HIV-1 use the human machinery to produce its RNA transcripts for protein expression and ultimately genomic replication. How viruses hijack cellular processes through interactions with host macromolecules is a fundamental question in biology and medicine.
Learn more about Structural Control of Human Co-factors for Retroviral Gene Expression