Pre-mRNA Splicing for Treatment of Human Disease
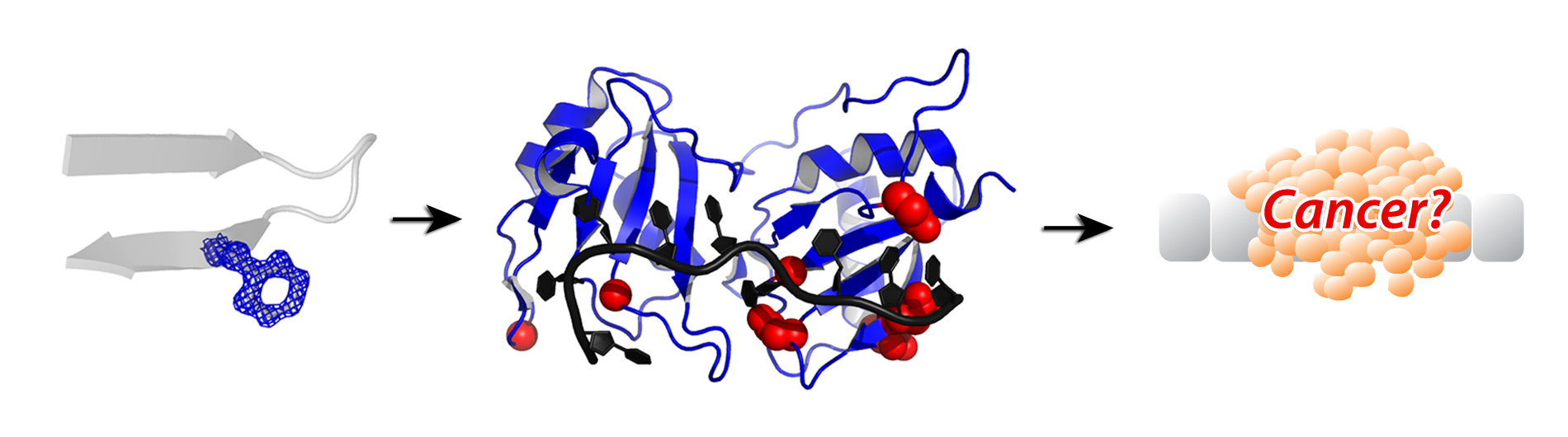
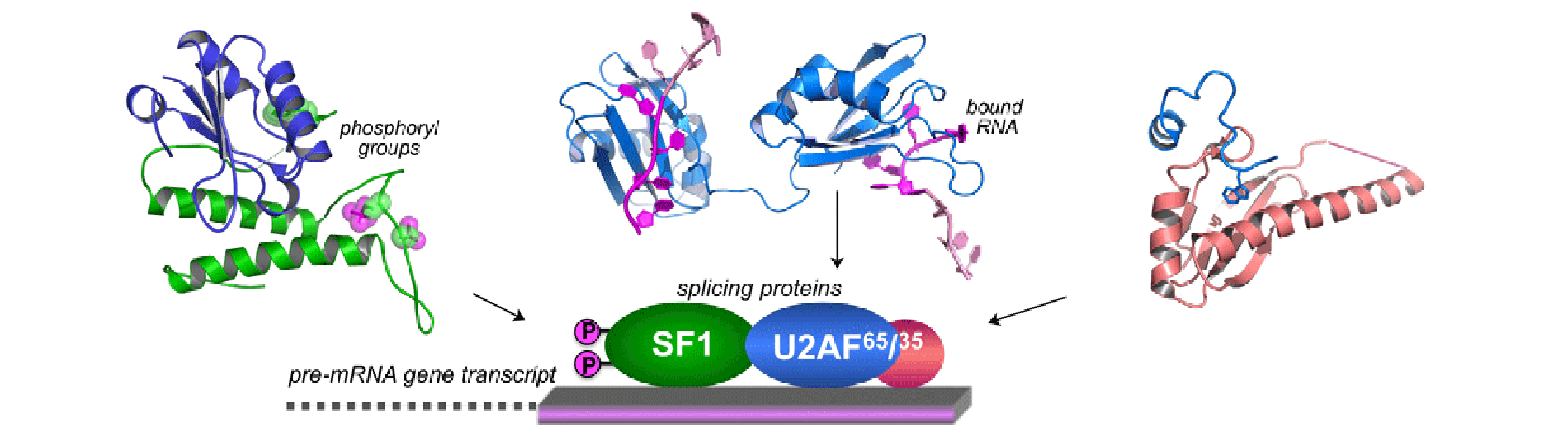
Human life depends on pre-mRNA splicing for cellular viability, differentiation and responses to changing physiology or environment. A major focus of my laboratory is to understand at a molecular level how the splicing machinery identifies sites for excision from gene transcript RNAs, which in turn changes the proteins produced. We have characterized the three-dimensional shapes of human splicing proteins recognizing one another and the gene transcript RNA at high resolution by X-ray crystallography (e.g. diagram above) complemented by molecular biology in human cells. Through this research, we identify a network of interactions responsible for recognizing human splice sites. The broader impact of this work for human disease is emphasized by the severe defects in pre-mRNA splicing that accompany most human hematologic malignancies and many metabolic disorders, as well as the dependence of HIV-1 and other complex retroviruses on RNA splicing for infectivity.

Clara L. Kielkopf, Ph.D.
Principal Investigator
- SF3B1 Phosphorylation Prompts U2AF2 Dissociation for Widespread Control of pre-mRNA Splicing.; bioRxiv : the preprint server for biology. 2026 Jan 20.
- Recurrent Neurodevelopmentally Associated Variants of the Pre-mRNA Splicing Factor U2AF2 Alter RNA Binding Affinities and Interactions.; Biochemistry. 2024 Oct 10.
- A UHM - ULM interface with unusual structural features contributes to U2AF2 and SF3B1 association for pre-mRNA splicing.; The Journal of biological chemistry. 2022 Jun 30.
- Pre-mRNA splicing factor U2AF2 recognizes distinct conformations of nucleotide variants at the center of the pre-mRNA splice site signal.; Nucleic acids research. 2022 May 07.
News
Affiliations
- Biochemistry & Biophysics
- Center for AIDS Research
- Center for RNA Biology: From Genome to Therapeutics
- James P. Wilmot Cancer Center
- NIH T32 Training Grant in Cellular, Biochemical & Molecular Sciences
- Biochemistry & Molecular Biology Ph.D. Program
- Biomedical Genetics and Genomics Ph.D. Program
- Biophysics, Structural & Computational Biology Ph.D. Program
- Wedekind Lab
Photos & Events
Contact Us
Kielkopf Lab
MC 3-8540
601 Elmwood Ave
Rochester, NY 14642