URMC / Labs / Lawrence Lab / Projects / Impact of environmental exposures during early life development on immune responses later in life, a
Impact of environmental exposures during early life development on immune responses later in life, and across generations
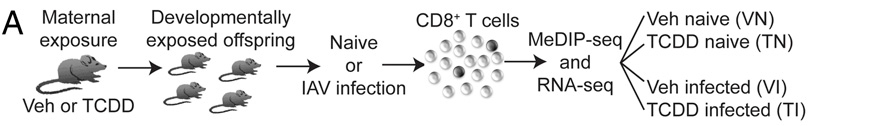
Early life exposures to environmental chemicals, and other differences in maternal and early life environment, can contribute to immune dysfunction later in life. Although these associations have been made, how exposure to environmental factors lead to changes later on is not fully understood.
« back to all projects