URMC / Labs / O'Connell Lab / Projects / Understanding the molecular mechanisms of differential microRNA-processing and its dysregulation in
Understanding the molecular mechanisms of differential microRNA-processing and its dysregulation in cancer
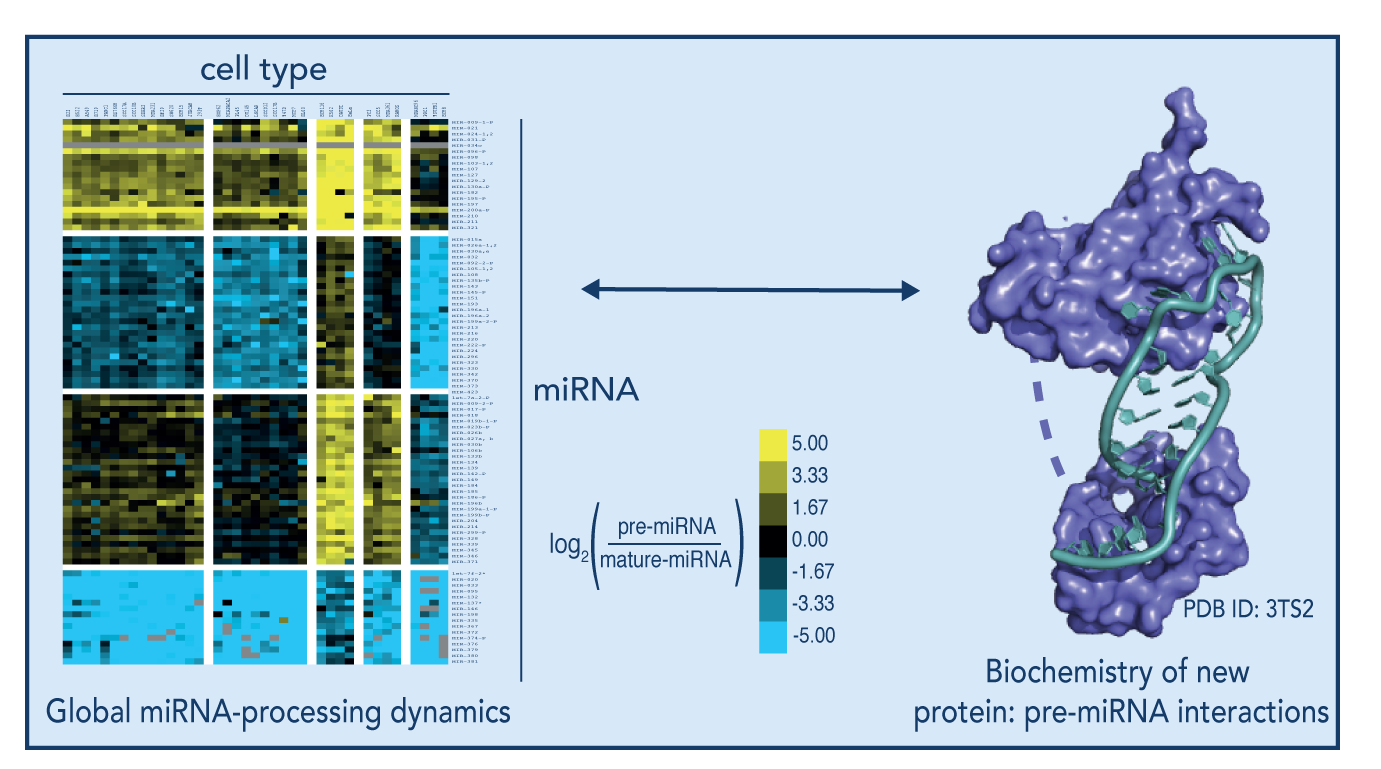
MicroRNAs (miRNAs) are small (21-24 nt) double-stranded RNA molecules that act to modulate eukaryotic gene expression through a process known as RNA interference (RNAi). miRNA precursors must undergo a number of processing steps after transcription to generate an active, mature miRNA. miRNA-processing appears to be regulated in both a miRNA-specific and cell-specific manner, suggesting that both precursor miRNA sequence elements and cell-specific factors contribute to the differential regulation of mature miRNA expression levels. It not known, however, the complete molecular mechanisms by which these miRNA precursors experience differential processing in different cell- and cancer- types.
The O’Connell lab is focused on understanding the mechanisms of differential miRNA processing by identifying and studying the protein partners responsible for the recognition of specific precursor miRNAs, with of the ultimate goal of understanding the mechanisms by which these protein partners affect miRNA processing in both normal and cancerous tissue. Using a CRISPR RNA-pull down technology, we have been able to identify a number of novel miRNA-binding proteins with specificity for particular miRNA precursors. We are following up on these leads with biochemical approaches to further elucidate the molecular determinants of these interactions, transcriptome-wide RNA-sequencing experiments to determine the global miRNA-binding landscape of these proteins, and the development of knockout (and/or mutant) cell-lines using CRISPR/Cas9 technology to understand the affect these novel miRNA-binding proteins have on miRNA processing in live cells.
Overall, the results of this work will yield a greater molecular understanding of how factors involved in regulating microRNA expression contribute to microRNA dysregulation in cancer, and will potentially provide new therapeutic routes for the treatment of this disease.